Description
T7 High Yield RNA Synthesis Kit optimizes the transcription reaction system. The kit can synthesize the single-stranded RNA efficiently by using T7 RNA polymerase, the linear double-stranded DNA with the T7 promoter sequence as the template, NTPs as the substrate to control the DNA sequence downstream of the promoter. During transcription, modified nucleotides can be added to the substrate to prepare biotin or dye-labeled RNA.
This kit can synthesize long transcripts and short transcripts, RNA can be produced 100-200 μg with 1 μg of DNA template input. The RNA synthesized by transcription can be used for various downstream applications, such as RNA structure and function research, RNase protection, probe hybridization, RNAi, microinjection, and in vitro translation.

Figure 1: In vitro RNA transcription process
Feature
- Up to 180 μg of RNA per reaction from 1 μg of the control template
- Optimized reaction system for the IVT process
- Decrease the dsRNA production
- Higher RNA integrity and purity
Application
- In vitro RNA synthesis
Components
| Components No. | Name | 10623ES50 (50 T) | 10623ES60 (100 T) | 10623ES70 (500 T) |
| 10623-A | T7 RNA Polymerase Mix | 100 μL | 200 μL | 1 mL |
| 10623-B | 10×Transcription Buffer | 100 μL | 200 μL | 1 mL |
| 10623-C | ATP (100mM) | 100 μL | 200 μL | 1 mL |
| 10623-D | CTP (100mM) | 100 μL | 200 μL | 1 mL |
| 10623-E | GTP (100mM) | 100 μL | 200 μL | 1 mL |
| 10623-F | UTP (100mM) | 100 μL | 200 μL | 1 mL |
| 10623-G | Control DNA Template (500ng/μL) | 10 μL | 20 μL | 100 μL |
Shipping and Storage
Dry ice transportation. Store at -15℃ ~ -25℃, valid for two years.
Figures
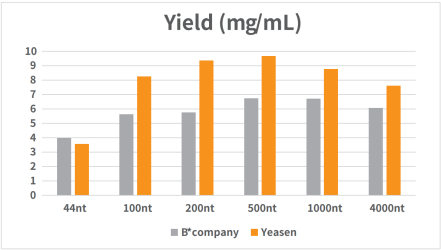
Figure 1. Standard RNA was synthesized in vitro using T7 RNA synthesis kit.
The reaction was incubated in a PCR instrument at 37℃ for 2h and then purified by magnetic beads (Cat#12602). The yield result was analyzed by NanoDrop spectrophotometer as shown in Figure 1.
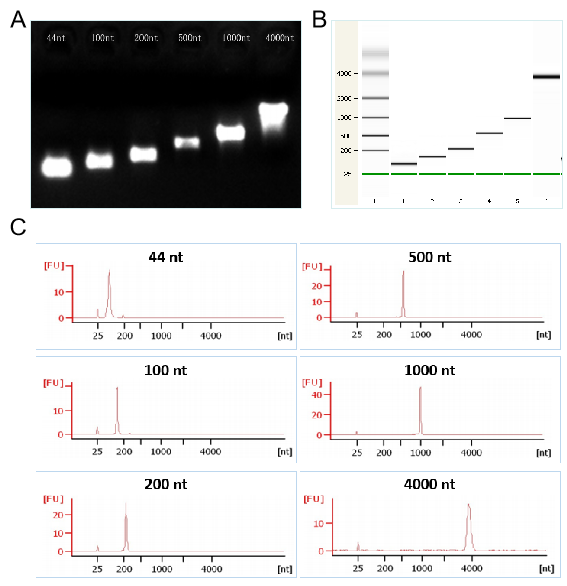
Figures 2. The transcription demonstration of different lengths of RNA by T7 kit respectively in the electrophoretogram (Figures 2A), the capillary electrophoresis diagram (Figures 2B), and the chromatogram (Figures 2C)
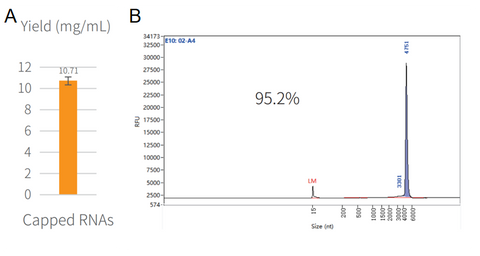
Figure 3. Synthesis of capped RNA in vitro.
The reaction was incubated in PCR instrument at 37℃ for 2h, and then purified by magnetic beads (Cat#12602). The yield result was assayed by NanoDrop spectrophotometer as shown in Figure 3A. The integrity result was analyzed by capillary electrophoresis as shown in Figure 3B.
1. Low transcript yield
The quality of the template is closely related to the yield. If the yield of the experimental group is significantly lower than the control group, the possible reasons are:
① the experimental template contains inhibitory components;
② The template has something wrong.
Suggestions:
① Re-purify the template;
② Determine the template quantification and its integrity;
③ Extend the reaction time;
④ Increase the amount of template input;
⑤ Try other promoters and RNA polymerases.
2. Low yield of short transcripts
A short transcription initiation fragment will inhibit the reaction. When the transcription product is less than 100 nt, extending the reaction time to 4-8 hs or increasing the amount of template to 2 μg will increase RNA yield.
3. RNA transcription length is greater than expected
If the electrophoresis shows that the product band is larger than the expected size, the possible reasons:
①The plasmid template may not be completely linearized;
②The 3' end of the sense strand has a prominent structure;
③The RNA has a secondary structure that is not completely denatured.
Suggestions:
①Check whether the template is completely linearized, and if necessary, perform additional linearization;
②Select a suitable restriction enzyme to avoid 3' overhangs, or use Klenow Fragment /T4 DNA polymerase to complete the transcription before proceeding;
③Use denatured gel to detect RNA products.
4. RNA transcription length is less than expected
If the electrophoresis shows that the product band is smaller than the expected size, the possible reasons:
①The template contains a termination sequence similar to T7 RNA polymerase;
②The GC content in the template is high.
Suggestions:
①Lower the reaction temperature (for example, 30°C). Sometimes lowering the temperature can increase the transcription length, but it will reduce the yield. Or try different RNA polymerases for transcription;
②If the template GC content is high, use 42℃ to transcript, or add SSB to increase the yield and transcription length.
5. Electrophoresis tailing of transcription products
There is a tailing phenomenon during electrophoresis.
Possible reasons:
①Contaminated by RNase during experimental operation;
②Contaminated DNA template by RNase.
Suggestions:
①Use RNase-free pipette tips and EP tubes, wear disposable latex gloves and masks, and all reagents are prepared with RNase free H2O.
②Re-purify the template DNA.
[1] Dong Z, Zheng N, Hu C, et al. Nosema bombycis microRNA-like RNA 8 (Nb-milR8) Increases Fungal Pathogenicity by Modulating BmPEX16 Gene Expression in Its Host, Bombyx mori. Microbiol Spectr. 2021;9(2):e0104821. doi:10.1128/Spectrum.01048-21(IF:7.171)
[2] Wang X, Tang S, Ye S, et al. Ultrasensitive quantitation of circulating miR-195-5p with triple strand displacement amplification cascade. Talanta. 2022;242:123300. doi:10.1016/j.talanta.2022.123300(IF:6.057)
FAQ
Q: What is the length of the Control DNA Template?
A: The size is 1142 base pairs.
Q: Does the T7 kit have a cap structure and polyA?
A: The polyA tail is introduced during the design of the template, while the addition of the cap is done separately.
Q: How can the IVT system be optimized?
A: ① Increase the input ratio of NTP and its analogues; ② Reasonable application of additives; ③ The reaction temperature can be set at 40 degrees.
The optimization of the IVT system can be achieved by adjusting the template, the amount of T7 enzyme used, and if it is co-transcription with cap addition, the ratio between the cap and NTP can be considered; for enzymatic cap addition, the conditions can be optimized in terms of the dosage of the cap-adding enzyme, 2-O-methyltransferase, and SAM. Regarding the template, apart from the requirement of complete linearization, there is another point, which is sequence design. For example, in the 5'UTR region, pay attention to uAUG and Kozak sequence; for CDS, consider the codon preference, GC3 content, and dinucleotide preference, etc.; 3'UTR: RBP and miRNA binding sites: and another point is the secondary structure of RNA. The stable secondary structure of the 5'UTR pre-sequence will reduce the expression of mRNA; the stable secondary structure of 3'UTR will increase the stability of mRNA and promote the expression of the encoded protein. For the recommended IVT system and more detailed tips, please refer to the mRNA in vitro synthesis experiment manual of Vivian Biotechnology.
Q: When the IVT reaction is scaled up from a 20 ul system to a 10 ml system, the yield decreases, and the yield per unit volume of the product drops by approximately 50%. Have you encountered this situation before? Do you have any suggestions?
A: When scaling up the 20uL system proportionally to 10mL, the yield should be approximately the same. Usually, we directly scale up from 20uL to 50mL, and the yield is generally at the level of 100ug/20uL. For the 50mL volume, we also use centrifuge tubes and a metal bath for measurement. For the 10mL volume, the amplification effect is generally better. In such cases, it is generally recommended to fine-tune the final concentrations of the enzyme and NTP before trying again.
Q: How much of the mRNA integrity can be achieved at present?
A: Regarding the re-submissions from our other clients, based on the CE results measured using Qsep100, the ideal situation is that the CE integrity can reach 88% to 92%; of course, achieving this level usually requires a series of optimizations.
The average yield is between 120-220 ug/ul (the yield is highly dependent on the sequence design). In the co-transcription method, the mRNA yield is 8 g/L. With process optimization, the yield will increase. Currently, the quality parameters of the available processes include observing the fluorescence value of GFP and measuring the cap addition rate using LC-MS.
Q: During your own testing process, do you use agarose gel electrophoresis to detect the mRNA products?
A: We can also conduct RNA electrophoresis. The main purpose is to briefly assess the purity of the products and check for any obvious aggregates, etc. If we are looking at the size of the RNA, during the electrophoresis process, we usually add a sample of a clearly defined RNA size as a reference.
Q: Will you conduct a test for the remaining Rnase in the DNA template?
A: Currently, the detection of the template mainly includes items such as sequence integrity, endotoxin, and residual host nucleic acids. There is no clear detection method for RNase yet; or you can refer to the detection method for residual RNase in our enzyme products, which is relatively simple and reliable.
Q: What is the mechanism for the generation of dsRNA during the IVT reaction?
A: There are several possible reasons for the generation of ds during the IVT process, mainly:
The T7 pol enzyme will use the product RNA as a template and continue to extend the RNA at its 3' end. The extended sequence will form complementary pairing with the 3' end sequence of the original RNA molecule, thereby creating a hairpin structure.
② The T7 pol enzyme can use RNA as a template to replicate RNA molecules, and synthesize the complementary strand.
(At present, there is no effective method for removing dsRNA. However, there is a patent report suggesting that a fiber column (sigma) could be tried.)
Q: Is there a feasible method to control the content of dsRNA in the products of in vitro transcription?
A: The suggestions regarding optimizing the IVT system and adjusting the reaction conditions to inhibit dsRNA mainly include the following points:
Reduce the concentration of magnesium ions; however, reducing the concentration of magnesium ions will also lead to a decrease in the production of RNA, and a balance point needs to be reached.
② Increase the reaction temperature of IVT to 50-56℃. This temperature can effectively reduce the content of dsRNA in the product; however, the requirements for equipment for high-temperature reactions will vary, and it may also increase the cost.
③ In the IVT reaction system, certain dispersants such as urea and formamide are added; our process research team has also conducted some experiments on this route, which can achieve the effect of inhibiting dsRNA. However, whether it is suitable for process scaling-up still needs further research.
Q: What are the reasons for the formation of precipitates during the IVT reaction process, and how can it be optimized?
A: Internal research has studied this precipitation. It is a polymer composed of magnesium ions, RNA, sodium ions, and magnesium pyrophosphate. There is a coagulation effect, and the aggregates keep growing in size (similar gel-like, flocculent, or particulate precipitations all follow a similar principle). The effects of adding water and adding EDTA are similar, which also supports this statement. Reasons: 1. If the concentration of Mg ions is too high, it will aggregate with pyrophosphate to form magnesium pyrophosphate precipitation. However, it has a coagulation effect, continuously encapsulating nucleic acids to form a gel-like substance, resulting in a decrease in yield. 2. A high RNA production level will also cause precipitation.
This kind of precipitation occurs by chance and is also closely related to the sequence design.
Solution: (1) Reduce the concentration of magnesium ions, but this will result in a decrease in yield. The amount of T7 enzyme needs to be increased and the reaction time (from 2 hours to 3-4 hours) needs to be extended; (2) The form of NTP (the presence of magnesium ions at a high concentration, such as 46 mM, does not select NTP, but when the magnesium ion concentration decreases, the yield of NTP (Tris salt form) does not change much, and the yield of the triNa form of NTP decreases significantly.
Q: What are the considerations for reactor selection during the process scale-up of mRNA production?
A: You could consider WAVE, Haodao Fu reaction vessels, and Sartorius. The main thing is to find bags that are compatible with the existing IVT system. For the stirring tank, the rotational speed of the propeller also needs to be optimized based on the actual situation, as it is related to the type and size of the blades. It is recommended to consult the equipment manufacturer.
Q: Do you recommend the two-step method or the one-step method? Why are most of them still using the two-step method at present?
A: In industry, the most commonly used method for enzymatic cap addition is to treat IVT products with vaccinia virus cap-adding enzyme, which can modify them into Cap 0 mRNA. The Cap 0 structure can be further modified into Cap 1 (m7GpppmN) under the action of 2'O-methyltransferase. With enzymatic cap addition, the cap addition efficiency can reach over 95%. The co-transcription cap addition method is simple to operate, but because GTP competes with the cap dimer, the cap addition rate of this method is a little lower; both methods have their own advantages and disadvantages.
Q: What is the output of IVT in vitro transcription?
A: Currently, our system uses a 20-ul reaction volume. The output of in vitro transcription is 100-200 ug. The internal amplification system with a 10 ml volume can reach 50 mg - 100 mg.
Q: When the reaction time exceeds 2 hours during in vitro transcription, will the completeness and yield decrease? How can the generated mRNA be ensured to be stable?
A: It is recommended that the entire reaction time be between 2 and 3 hours. Our company has verified a maximum reaction time of 3 hours, and our cooperative clients have also verified a reaction time of 4 hours, and the results were normal.
Q: Could you please assist in designing the DNA template sequence for in vitro mRNA transcription (mainly for the application in COVID-19 vaccines)?
A: No related services are provided. Currently, the main reference for the COVID-19 vaccine is still the publicly available vaccine sequences from external sources, namely BioNtech and Moderna. The main original sequences are generally reliable, except for the incomplete polyA tail; the earlier COVID-19 vaccines developed in China are mostly based on the RBD protein sequences. We haven't obtained the specific sequences for this part; basically, these are the core technologies of each research team.
Q: What methods and instruments are used for the detection of the cap rate and tailing effect?
A: The capillary column used for adding the cap is from the Agilent series (6230B TOF), while the tailing column used is from Thermo's QE or Waters' Rda.
Q: Do you use tail-adding enzymes when adding the A tail? How many base pairs do you generally recommend for the connection?
A: This reagent kit does not contain the component for poly A tail. In the template preparation stage, the A tail can be attached to the plasmid, and the length is usually between 20 and 200.
Q: In the product purification process, what is the maximum speed for centrifugation? What is the corresponding rotational speed?
A: Any power rating above 13000 rpm is acceptable.
Q: Why did this B component of the Buffer form a precipitate? How should it be dealt with?
A: The precipitate is spermidine. This is a normal phenomenon and it can be dissolved by hand.
Related blog:
Payment & Security
Your payment information is processed securely. We do not store credit card details nor have access to your credit card information.
You may also like
Inquiry
FAQ
The product is for research purposes only and is not intended for therapeutic or diagnostic use in humans or animals. Products and content are protected by patents, trademarks, and copyrights owned by Yeasen Biotechnology. Trademark symbols indicate the country of origin, not necessarily registration in all regions.
Certain applications may require additional third-party intellectual property rights.
Yeasen is dedicated to ethical science, believing our research should address critical questions while ensuring safety and ethical standards.

